New Hope for Biocomputers: Parallel Network-Based Computation
Download this article in PDF format.
Through the Bio4Comp multidisciplinary project, researchers from different fields of expertise (e.g., mathematics, biology, engineering, and computation) plan to build a network-based biocomputer that will be competitive with other alternative computing approaches. Several types of biological computation already exist, such as DNA and quantum computation. However, they’re not scalable and are considered impractical from a fabrication and operational perspective.
It has been shown in practice, though, that network-based biocomputers are self-assembling and they can be used in large numbers. Therefore, they do hold the potential of being scalable through the use of molecular motors.
The idea of biological computers was born after the creation of nanobiotechnology, which made it possible to engineer biomolecular systems that can interact with the same functionality of a computer. The first DNA computer was invented by Leonard Adleman in 1994. In 2013, Stanford University invented the biological transistor, which was made from genetic material (DNA and RNA) in place of gears or electrons. The team from Stanford called its biological transistor the “transcriptor."
Now, Bio4Comp—a collaborative effort between Lund University, B CUBE-Center for Molecular Bioengineering, Linnaeus University, Molecular Sens Ltd., Bar Ilan University, and Fraunhofer-Gesellschaft—will focus on scaling up biological computers by using a parallel-computation system. It will include biomolecular motors as computing units, where biomolecular machines act as adding machines.
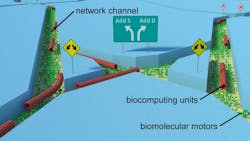
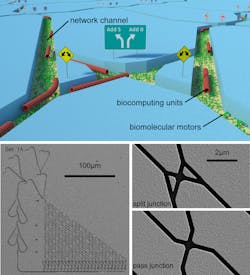
Filaments in the network travel very fast while following junctions (split and pass junctions) that lets them go stray, or turn right or left, allowing the motors to solve problems. (Source: Fraunhofer Institutes ISC and ENAS)
These biomolecular machines, each only a few billionth of a meter (nanometers) in size, can solve problems by moving themselves through a nanofabricated network of channels designed to represent a mathematical algorithm (see figure). Among the benefits of molecular motors, which have already been demonstrated, are:
• They move at a speed of 10 µm/s on the surface.
• Because they’re self-assembling, they can be used in large numbers. As a result, their computing power adds up quickly.
• They don’t need an external power supply. They can move based on a chemical added to the solution to propel them.
They also are extremely energy-efficient. The biomolecular motors will consume less energy while still having the ability to cool processors without obstructing the development of more powerful computers. Thus, they will be able to solve problems where many solutions need to be explored simultaneously. "One of the most exciting aspects of network-based computing with molecular motors is that it needs a hundred to thousand times less energy than electronic computers," says Hiener Linke, Project Coordinator at Bio4Comp.
“Practically all really interesting mathematical problems of our time cannot be computed efficiently with our current computer technology,” adds Dan V. Nicolau, Ph.D. M.D., from the U.K.-based enterprise Molecular Sense. He was one of the people who originally thought of using biomolecular motors as computers.
Nicolau was a collaborator in a previous publication from the Proceedings of the National Academy of Sciences (PNAS), which demonstrated the foundations of an alternative parallel-computation system. In that system, a given combinatorial problem is encoded into a graphical, modular network that’s embedded in a nanofabricated planar device using molecular-motor-propelled agents to solve a mathematical problem. This approach used orders of magnitude less energy than conventional computers, hence addressing issues related to power consumption and heat dissipation.
If biological computers can be successfully scaled, solutions for the world of computing will be limitless. Maybe one day, when Moore’s law completely stalls, parallel-network biocomputing will take over power electronics.
About the Author
Maria Guerra
Power/Analog Editor
Maria Guerra is the Power/Analog Editor for Electronic Design. She is an Electrical Engineer with an MSEE from NYU Tandon School of Engineering. She has a very solid engineering background and extensive experience with technical documentation and writing. Before joining Electronic Design, she was an Electrical Engineer for Kellogg, Brown & Root Ltd (London. U.K.). During her years in the Oil and Gas Industry she was involved in a range of projects for both offshore and onshore designs. Her technical and soft skills bring a practical, hands-on approach to the Electronic Design team.